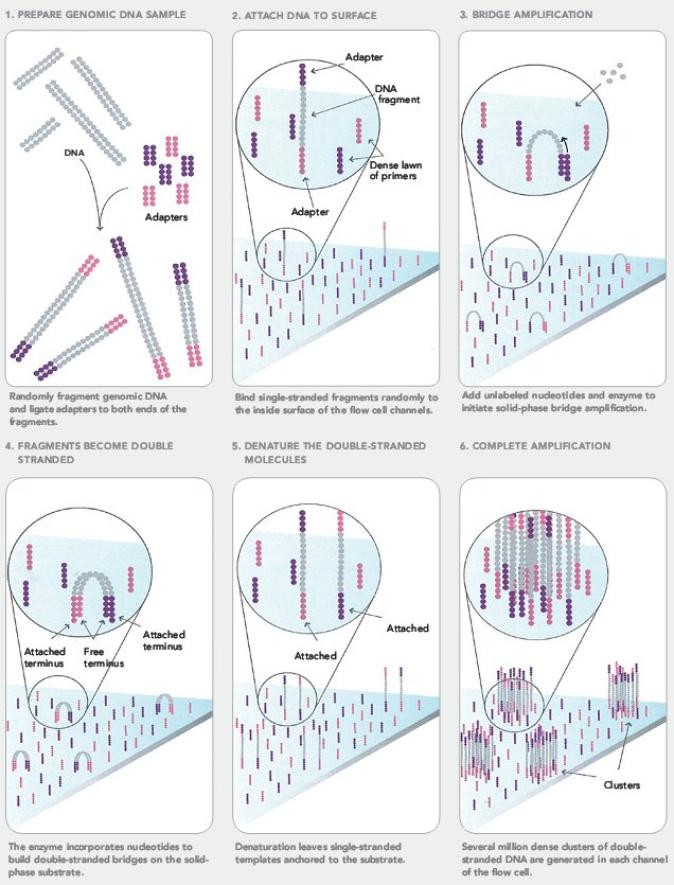
We will now take a closer look at the Illumina sequencing method, which uses a technique called reversible termination sequencing:
|
Click to enlarge.
In this video, pay close attention to how bridge amplification works. |
|
3. Next,
the extra bases and DNA polymerase are "washed" off. Instead of
needing a capillary tube for electrophoresis, cameras can simply record
the color of the RT-base added to the end in the "scanning"
phase, since each RT-base has its own color: blue for cytosine RT-bases,
green for thymine RT-bases, yellow for guanine RT-bases, and red for
adenine RT-bases.
4. The dye attachment at the end of the RT-base is subsequently cut off so that the next base to be sequenced is exposed, and DNA polymerase and bases are added again, going back to step two.
5. This "wash-and-scan" cycle repeats until the entire DNA fragment has been sequenced; it is done simultaneously with millions of DNA fragments at a time and can sequence a complete human genome in less than a week. (31) (38)
4. The dye attachment at the end of the RT-base is subsequently cut off so that the next base to be sequenced is exposed, and DNA polymerase and bases are added again, going back to step two.
5. This "wash-and-scan" cycle repeats until the entire DNA fragment has been sequenced; it is done simultaneously with millions of DNA fragments at a time and can sequence a complete human genome in less than a week. (31) (38)